Ultraviolet Microscope To Dramatically Speed Up Lab Tests
By Jof Enriquez,
Follow me on Twitter @jofenriq
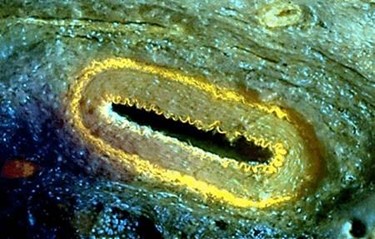
A microscope using ultraviolet (UV) light to provide high-resolution histological images promises to significantly shorten the amount of time diagnostic tests are processed.
Commonly, lab technicians and researchers using bright-field (transmission) clinical microscopes have to slice tissue 4-6 micrometers thin onto glass slides, which takes hours to days to prepare. Researchers at University of California, Davis, propose a new type of microscopy to quicken the process to mere minutes.
Their approach, called microscopy with ultraviolet surface excitation (MUSE), uses UV light at wavelengths below 300 nanometers to penetrate tissue samples by only a few microns below the surface (roughly the same thickness of a typical histology slide), without having to destructively section and stain tissue slices.
“MUSE eliminates any need for conventional tissue processing with formalin fixation, paraffin embedding or thin-sectioning,” said Richard Levenson, professor and vice chair for strategic technologies in the Department of Pathology and Laboratory Medicine at UC Davis.
“It doesn’t require lasers, confocal, multiphoton or optical coherence tomography instrumentation, and the simple technology makes it well suited for deployment wherever biopsies are obtained and evaluated," he added.
The optical system comprises one or more widely available UV light-emitting diodes (LEDs), and a UV-compatible sample stage, complemented by conventional microscope components.
As explained in Nature Bioemedical Engineering:
UV light at the 280 nanometer spectral range (maximum output of 0.9mW per LED) illuminates an approximately 1 square millimeter area of the specimen at an oblique excitation angle, yet restricted to regions very close to the surface (a few nanometers deep), provides sharp, high-contrast images. The excitation light at the sub-300 nm spectral region elicits bright emission from tissue specimens stained with conventional fluorescent dyes. These stains emit photons in visible-band signals that are captured using simple-to-operate and inexpensive conventional glass-based microscope optics, and either grayscale or color cameras.
Since histologists and pathologists are used to viewing bright-field, H&E (haemotoxylin and eosin) stained specimens, MUSE images are converted by a Python-language utility run on a graphics processing unit, in real time, into images that are near identical to the more familiar and authentic H&E appearance.
Because conventional tissue sectioning is not required to elicit these high-resolution images, it will take a few minutes, rather than hours or days under current methods, to stain fresh or fixed tissues. The non-destructive nature of MUSE can be particularly helpful in diagnosing cancer. Since tissue samples are left intact, biopsy specimens that have been imaged can then be forwarded to other laboratory tests, such as downstream molecular assays (including fluorescence in situ hybridization and RNA sequencing).
“It has become increasingly important to submit relevant portion of often tiny tissue samples for DNA and other molecular functional tests,” Levenson said. “Making sure that the submitted material actually contains tumor in sufficient quantity is not always easy and sometimes just preparing conventional microscope slices can consume most of or even all of small specimens. MUSE is important because it quickly provides images from fresh tissue without exhausting the sample.”
As a fluorescence-based, slide-free optical imaging system, MUSE has the potential to significantly speed up diagnostic tests, alleviate patient anxiety over waiting, and cut technical costs. In the United States alone the current cost of preparing hundreds of millions of slides is estimated at billions of dollars, according to the researchers.
In addition, as a microscopy technique alone, MUSE can benefit areas such as basic biology, microanatomy, toxicology, agriculture, and education, they added.
Like the UC Davis researchers, other scientists are keen on developing optical alternatives to acquiring cellular-scale images without sectioning tissue samples required by current histological methods. These new approaches include optical coherence tomography, structured illumination, conventional reflectance and fluorescence confocal microscopy, and multi-photon imaging, among others in development.
